Deadly Disease Outbreak Contained by Genome Sequencing
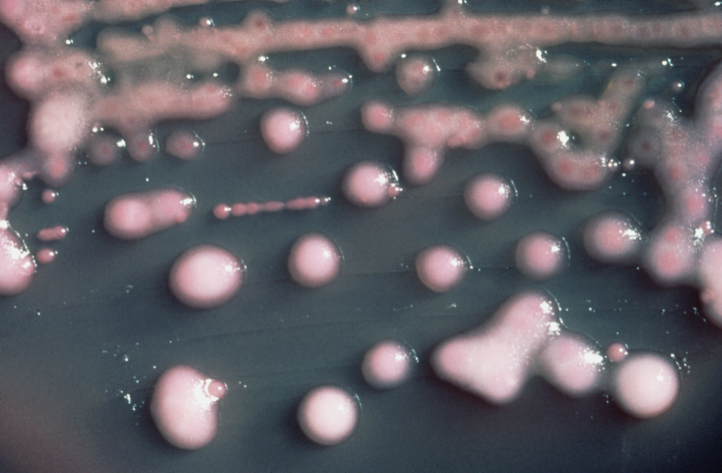
Genome sequencing helped researchers track a deadly infection that broke out in a research hospital last year. The infection was caused by multiple-drug resistant bacteria, K. pneumoniae. The bacteria infected 18 people, of whom 11 died.
The National Human Genome Research Institute (NHGRI) along with staff from Clinical Center of the National Institutes of Health in Bethesda, Md. then set off to get the disease under control. They used genome sequencing to track the spread of infection in real-time. They also zeroed-in on the very first patient who got the infection.
"Infectious outbreaks happen in every hospital in the world, afflicting millions of patients each year in the United States alone. By marshaling the ability to sequence bacterial genomes in real time to accurately trace the bacteria as it spread among our Clinical Center patients, our researchers successfully elucidated what happened, which in turn has taught us some important lessons. This study gives us a glimpse of how genomic technologies will alter our approach to microbial epidemics in the future," said NHGRI Director Eric D. Green, MD, PhD.
"For decades, we used pulsed-field gel electrophoresis to differentiate bacterial strains," said Tara N. Palmore, MD, the NIH Clinical Center's deputy hospital epidemiologist who led the outbreak investigation.
The outbreak began in June 2011 when a patient from a New York City hospital was sent to the NIH Clinical Center. The 43 year old patient had a medical history of drug-resistant infections. She was immediately isolated from the rest of the patients and treated separately. Despite all the precautions taken by the NIH, other patients in the hospital began getting infected from the deadly strain as many were on immunosuppressant drugs.
When the outbreak was reported, hospital team sent samples of bacteria taken from infected persons to the NIH Intramural Sequencing Center (NISC). It was at NISC where the DNA of the bacteria was sequenced. A team, led by Julie Segre, PhD, who is an NHGRI senior investigator, analyzed the genome sequence.
The researchers found that the outbreak was caused by a single patient from New York or that there was just one source of infection. The genomic sequence data was then combined to traditional epidemiological tracking methods. Researchers found that he first patient had infected two more patients thus creating two major clusters of infected patients.
"We were already trying to develop clinical molecular diagnostics tools. We thought we could use genome sequencing to tell whether the K. pneumoniae from the first patient was the same strain as the one that infected the second patient," Dr. Segre said.
The hospital staff was able to contain the spread of infection by limiting exposure of the strain to other patients and staff members. The first patient of the infection was treated with a potent antibiotic drug called colistin which is considered a last resort for fighting an infection.
"Genome sequencing and analysis is our best hope for anticipating and outpacing the pathogenic evolution of infectious agents. Though our practice of genomics did not change the way patients were treated in this outbreak, it did change the way the hospital practiced infection control," said Segre.
The genome sequencing efforts were published in the journal Science Translational Medicine.
Published by Medicaldaily.com